NOT FOR USE IN DIAGNOSTIC PROCEDURES (EXCEPT AS SPECIFICALLY NOTED). Our mission is to develop high-quality innovative tools and services to accelerate discovery.įOR RESEARCH USE ONLY. As a member of the Takara Bio Group, Takara Bio USA is part of a company that holds a leadership position in the global market and is committed to improving the human condition through biotechnology. Design the overlapping primers for the amplification of the fragment B following the same rules. provides kits, reagents, instruments, and services that help researchers explore questions about gene discovery, regulation, and function. Check secondary structures of the generated primers by online oligo analyzers (see Note 9) strong intramolecular secondary structure within the primers can inhibit the PCR reaction and give low yields of product. To add the primer at the selected location, go to Primers Add Primer. If you click the mouse and drag to select, the melting temperature of a corresponding primer will be shown. 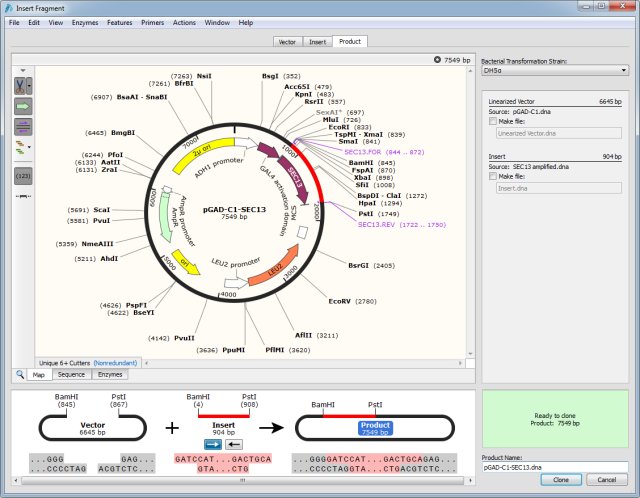
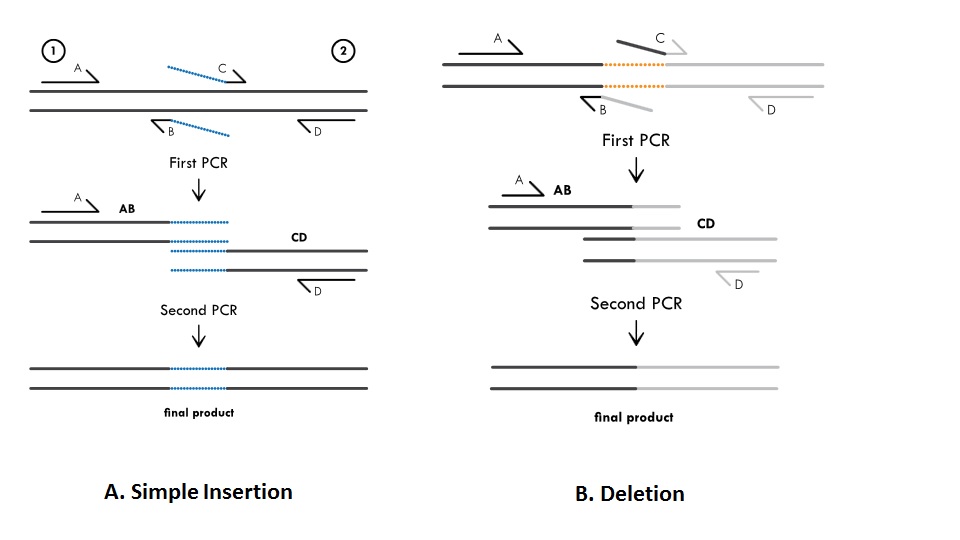
Alternatively, you can begin by selecting the desired binding site on your target sequence. Click Assemble to run the CAP3 assembler. Option 2: Create a Primer Based on a Target Sequence. Provide a name for your assembly and set the Save destination for the assembly. © 2018 Cold Spring Harbor Laboratory Press.Takara Bio USA, Inc. Ensure the option to 'Trim low-quality ends of sequences before running CAP3' option is checked. Learn how to annotate your sequences easily using the enzymes, features, and primers menus. Here are the steps to follow to design primers to amplify exon 15 from human BRCA1: 1. This Getting Started video introduces SnapGene annotations. This method can also be used to generate deletions, which is discussed in the latter part of the protocol. In this article, we will show you how to use NCBI Reference Sequences and Primer-BLAST, NCBI’s primer designer and specificity checker, to design a pair of primers that will amplify a single exon (exon 15) of the human breast cancer 1 (BRCA1) gene. In the secondary PCR, the insertion fragment is recombined with two flanking fragments derived from the original template. To generate an insertion mutation, first prepare an insertion fragment and two flanking fragments by PCR. Using this method, one can essentially make insertions and deletions of any size at any position within a given DNA sequence. Create a Primer (SnapGene 6.2 and earlier) Edit Multiple Primers. This protocol describes an overlap extension PCR mutagenesis method that is more versatile than its predecessors. For example, traditional protocols require that all sequence alterations be embedded within the primer itself, which makes it difficult to make insertions >30 nt. overlapping primers for the assembly of Fragment 1 and Fragment 2 into the. Review and if required, reverse the orientation of the insert. SnapGene (or an equivalent sequence viewer) and using the variant. Alternatively, use the drop-down menus or type the enzyme names in the boxes at the upper right. Traditional overlap extension PCR mutagenesis protocols remain limited in several critical ways, especially when it comes to generating insertions and deletions. To specify the fragment to be inserted: Click on the first enzyme site, then either Shift-click on the second enzyme site or drag to the second enzyme site. Gibson Assembly: This technique joins PCR amplified DNA segments to create overlapping ends and inserts fragments into a plasmid without the. Just select the DNA fragments to fuse, and SnapGene will design the primers. Unlike methods of random mutagenesis, directed mutagenesis requires that the researcher already have a specific mutation in mind to implement. For the In-Fusion reaction, a linearized vector is mixed with one or more PCR products with overlapping ends. In-Fusion Cloning: SnapGene is the first software to simulate Clontech’s versatile method for creating seamless gene fusions. Because of its versatility, the method has become widely used. It requires relatively little preparation compared with other mutagenesis methods and does not require the use of restriction enzymes. Overlap extension polymerase chain reaction (PCR) mutagenesis can be used for the generation of a specific point mutation, insertion, or deletion within a particular DNA sequence of interest.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
